Dragon 5.3 has been greatly improved, especially with regard to precision in descriptor calculation. Bugs have been fixed and corrections to some descriptor algorithms have been performed in order to get sounder values. Some new descriptors have been added, increasing the number of calculated molecular descriptors to 1664.
A new version working in Linux system has been released with the name dragonX. At the moment, this version can only work in background mode by a command line. dragonX has no restriction on the number of processed molecules and on the molecule size; so far, it has been tested up to 250,251 molecules (NCI database) and 700 atoms per molecule.
New atom types have been added to the set of atom types recognized by Dragon: As, Se, Te, Ga, In, Tl, Pb.
Most of the common charged compounds are now correctly processed. Algorithms for descriptor calculation have been changed to account for atom formal charges.
The maximum atom connectivity (i.e., the number of bonds) has been increased up to 10.
A new algorithm for SMILES notation reading has been implemented. It is based on the basic rules proposed by Weininger (Weininger D., J.Chem.Inf.Comput.Sci. 1988, 28, 31-36). Most of the previously rejected SMILES are now correctly processed.
Major effort has been expended in order to obtain a standard internal representation of molecules regardless of the format of the input files. The aim was to obtain unique descriptor values for a given molecule starting from different representations. It's well-known that the common MDL file represents the molecule by means of an aliphatic structure (i.e., a Kekulè-like structure), while HyperChem files and SMILES notations can give different molecule representations according to how the user has defined conjugated systems and aromaticity.
In the Dragon molecule representation, all bonds belonging to aromatic rings are assigned a conventional bond order of 1.5 while bonds belonging to non-aromatic conjugated systems are alternating single and double bonds. Delocalized bonds, such as in the nitro group, are represented as covalent bonds to both uncharged oxygens (N=O). Aromaticity detection is performed by an internal algorithm and thus it is independent of the aromatic bonds specified by the user.
The user is free to choose whether to use the standard Dragon molecule representation or keep the one defined in the input file, by the option 'User defined bond orders' in the 'Select files' menu. Note that this option is available only for Dragon run in stand-alone mode. When Dragon is running in background mode the bond orders are always recalculated except when the molecules are entered in the H-depleted form.
In the algorithm for MDL file reading a check has been introduced for the detection of molecules with radicals. These molecules, characterized by one or more value of 4 in the charge field, are rejected by Dragon.
The option 2D-structures in the 'Select files' menu has been included. By checking this option, calculation of 3D descriptors will be skipped. We would like to remind the user that 3D descriptors may be even calculated for 2D structures; however, calculation may fail whenever a molecule contains two atoms with similar coordinates. In effect, in this case the geometrical distance between the two atoms is near 0.
The script file used to run Dragon in background mode has been changed accordingly. The record No. 22 related to the input file has got a new field -d, which is optional:
/fm inputfilename.txt –fn –imolID [-hb] [-typed]
The variable type can take value 2, if the entered molecules are 2D structures, or value 3, if the molecules are 3D. If the field -d is not specified in the script, all the structures are supposed to be 3D.
The valence vertex degree formula has been changed to account for formal charges. The valence vertex degree of an atom is now calculated by subtracting both the H-bonds and the formal charge to the number of valence electrons.
To generate several output files, one for each selected descriptor block, the user previously had to specify one output file name with extension .bnn; now, the file name is allowed to have any extension but it must include the tag 'bnn', which can be placed in any position within the file name.
The nAB descriptor (number of conjugated bonds) has been redefined as the number of aromatic bonds. While previously it counted all the edges in the molecule with conventional bond order equal to 1.5, assigned to bonds in any conjugated system such as C = C - C = C and C = N - C = C, now it only counts all the bonds belonging to aromatic rings.
The number of rotatable bonds (RBN) is now calculated according to the definition given in F. Veber et al., J.Med.Chem. 2002, 45, 2615-2623. Rotatable bonds are defined as any single bond, not in a ring, bound to a nonterminal heavy atom. Excluded from the count are amide C - N bonds because of their high rotational energy barrier.
The calculation of topological polar surface area (TPSA), based on the atom-based method of Ertl, Rohde, and Selzer, has been greatly improved, especially with regard to the recognition of atoms belonging to aromatic rings. In fact, according to the TPSA method a ring is aromatic if it doesn't include any sp3 carbon. Moreover, two different TPSA values are now calculated, namely TPSA(NO) and TPSA(tot), the first being derived only from polar fragments with nitrogen and oxygen and the second from polar fragments with nitrogen and oxygen plus "slightly polar" fragments containing phosphorus and sulphur.
The list and definitions of functional groups has been revised. Some new groups have been added such as thioureas, carbonates, anhydrides, hydrazones, amidines, guanidines and several heterocycles. The hydrogen bond donors and acceptors are calculated differently. The number of intramolecular H bonds has been also added. A detailed list of the functional groups with rules for their identification is given in the molecular descriptor theory section.
The number of donor atoms for H-bonds (nHDon) is now calculated by adding together the hydrogens bonded to any nitrogen and oxygen.
The number of acceptor atoms for H-bonds (nHAcc) is now calculated by adding together any nitrogen, oxygen and fluorine, excluding N with formal positive charge, higher oxidation states and pyrrolyl form of nitrogen.
The number of intramolecular H bonds (nHBonds) is a new molecular descriptor, which adds up any atom pairs Y1Y2 in the molecule so that Y1 can be B, N, O, Al, P, S with at least one bonded hydrogen and Y2 can be N, O, F. Moreover, to have an intramolecular H bond the topological distance between Y1 and Y2 must be 3 or 4 and the geometrical distance between Y2 and H bonded to Y1 must be in the range (1; 2,7). Note that to calculate this descriptor, which is based on geometrical distances between atoms, it's necessary to select the geometrical descriptor block along with the functional groups (this only holds for Dragon for Windows).
In the block of charge descriptors, the maximum negative charge and the total negative charge are now reported with the minus sign and no longer with the absolute value. Moreover, charge descriptor calculation has been also extended to MDL molecule files, where atomic formal charges can be specified. However, we would like to remind the user that most of these descriptors are meaningful only if derived from atomic partial charges.
A new algorithm for the calculation of neighbourhood symmetry indices (ICs, TICs, BICs, SICs) has been implemented. Moreover, for small molecules, some higher-order indices may now have missing value because atom equivalence with respect to higher-order neighbourhood cannot be determined.
The E-state topological parameter (TIE) has been modified with respect to the original one [A. Voelkel, Computers Chem. 1994, 18, 1-4] in order to obtain more well-founded values for all molecules. It is now calculated as follows:
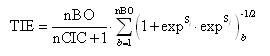
where nBO is the number of non-H bonds, nCIC the number of rings in the molecule, Si and Sj the electrotopological state indices for the two atoms incident to the bth bond.
The Ghose-Crippen logP and molar refractivity based on a group contribution method have been improved even if some ambiguities still remain in recognition of some atom-types. The atom-type fragment counts have been changed accordingly.
The Moriguchi logP algorithm has been greatly improved. Specifically, the PRX variable has been given a new interpretation on the basis of the rules and examples given in the original papers of Moriguchi; the RNG variable is now calculated by considering as aromatic any ring without sp3-hybridized carbon atoms; the AMP variable now takes value of 1 also for any hydrocarbon chain connecting -COOH and -NH2 groups. All the MlogP values published in the Moriguchi's original papers have been correctly reproduced.
The AlogP and MlogP models implemented in Dragon have been evaluated on a set of 2648 compounds with experimental logP taken from the NCI Open Database. The resulted correlation coefficient r is 0.915 for AlogP and 0.915 for MlogP.
Parameters on which the Lipinski's Alert Index is based have been revised. While the H-bond donor variable is calculated by adding all of the OHs and NHs bonds in the same way as the Dragon HDon variable, the H-bond acceptor variable is calculated by adding all of the Os and Ns as proposed in the original Lipinski's paper. For what is concerned with the MlogP, we still prefer using the algorithm implemeted in Dragon (now greatly improved) instead of the MlogP calculation rules proposed by Lipinski, which show some ambiguity and partly differ from the original ones of Moriguchi.
The Ghose-Viswanadhan-Wendoloski drug-like indices, previously only based on the range of some pysicochemical properties, now also account for the presence of specific structural features in the molecule.